MicroRNAs and autism spectrum disorder: small tools for a complex disorder
- Inicio
- Comité Editorial
- Lineamientos
- Carta de Cesión de Derechos
- Información Legal
- Acerca de la Revista
- Bases de Datos
- Contacto
- ISSN 2007-3054
- Centro de Investigaciones Cerebrales
Universidad Veracruzana
Jesus Antonio Lara-Reyes1; Alejandra Llanes-Duran1; Gonzalo Emiliano Aranda-Abreu2; María Elena Hernández-Aguilar2; Fausto Rojas-Durán*2
1Doctorado en Investigaciones Cerebrales, Universidad Veracruzana, campus Xalapa
2Centro de Investigaciones Cerebrales, Universidad Veracruzana, Xalapa, Ver. México
Abstract
Resumen
Introduction
Autism spectrum disorder
ASD genetics
miRNAs
miRNAs and ASD
miRNAs as biomarkers and their therapeutic potential in ASD
Conclusion
Conflict of interests
Acknowledgements
References
Mail
Abstract: The autism spectrum disorder (ASD) includes a complex group of neural development disorders that are characterized by deficiencies in social interaction, communication, and motor skills of individuals. ASD has an impact on public health, so it would be very useful to have biomarkers that allow early identification of the disorder. In this sense, microRNAs (miRNAs), which are regulators of a wide variety of cellular functions and whose alterations in their expression have been observed in individuals with ASD, are beginning to be considered as potential targets for the development of diagnostic and therapeutic strategies for these disorders. This review focuses on some studies about the participation of miRNAs in ASD in animal models, as well as in humans, and makes an approach to their possible use as biomarkers.
Keywords: MicroRNAs, autism spectrum disorder, biomarker.
El trastorno del espectro autista (TEA) abarca un grupo complejo de trastornos del desarrollo neural que se caracterizan por presentar deficiencias en la interacción social, la comunicación y motricidad del individuo. El TEA tiene un impacto en la salud pública, por lo que sería de gran utilidad contar con biomarcadores que permitan la identificación temprana del trastorno. En este sentido, los microRNAs (miRNAs), que son reguladores de una gran variedad de funciones celulares y cuyas alteraciones en su expresión han sido observadas en individuos con TEA, se comienzan a considerar como blancos potenciales para el desarrollo de estrategias de diagnóstico y terapéuticas para estos trastornos. Esta revisión se enfoca en algunos estudios sobre la participación de miRNAs en el TEA, en modelos animales y humanos, y hace una aproximación a su posible uso como biomarcadores.
Palabras clave: MicroRNAs, trastorno del espectro autista, biomarcadores.
The miRNAs belong to a class of endogenous small non coding RNA that play a crucial role in the post-transcriptional regulation of gene expression,1 therefore they are regulator of several cellular activities and have been related to many diseases, one of them ASD.2 The ASD encompasses a complex group of neurodevelopmental diseases characterized by deficiencies in the individual´s social interaction, communication, and motor skills of the individual.3 Currently, there is still a lack of precise laboratory tests for the detection of ASD with specific observation of behavioural changes being the conclusive diagnosis.4 It is consider that its origin is both genetic and environmental, and it is very complex, with the participation of multiple genes, which in turn can be regulated or modified epigenetically.3,5 In this sense, it is where miRNAs become important, of which they have been found to have important roles in the different stages of brain function.6 This review addresses the relationship of miRNA with some aspects of ASD, emphasising their usefulness as biomarkers.
ASD is a complex group of disorders of neural development that have as characteristic deficiencies in social and emotional interaction (isolation), and communication of the individual (language disorders), as well as repetitive and stereotyped behaviours (motor disorders).1,5 It has been observed that it occurs more in boys than in girls in 3:1 ratio. ASD is expressed during early childhood, between 2 and 3 years approximately, and accompanied by cognitive, learning, attention, and sensory processing dysfunctions. Its clinical manifestations are over the years, which can be attributed to the child´s growth, brain development, therapies used, and environmental factors.3,8
Although ASD is considered to be caused by a combination of genetic and environmental factors, its origin remains unknown. It has been proposed to divide ASD into primary and secondary. In primary type cases, the data suggests a multifactorial etiology in which genetics is essential but not unique, and between 70 to 90 % of ASD cases fit with this type.5 Those of the secondary type, unlike those of the primary type, can be associated with a known cause, such as the use of certain drugs during pregnancy, intrauterine rubella, fragile X syndrome and tuberous sclerosis, the latter being the most common, and only represent between 10 and 30 % of all cases.5
It has been estimated that 1 of 68 children present ASD in the U.S. and it could occur in all racial, ethnical and socioeconomic group. Around the world the prevalence of this condition is around 1 and 2%.9 Mexico has a lack of data at national level about the prevalence of ASD. In a state-level study, it was estimated that 1 out of every 115 children has this condition,10 however, the amount of cases with this condition could be more across the country, not just because of the absence of data information, but also unawareness of the condition by family members, caregivers and teachers, in addition to the subjectivity of the behavioural tests carried out.11 What leads us in to the necessity to find reliable and easy to detect biomarkers to confirm the diagnosis. Given the above it must be recognized that ASD is not only a public health issue but also a problem that impacts the social environment of the child and a late diagnosis will limit his development in the society.
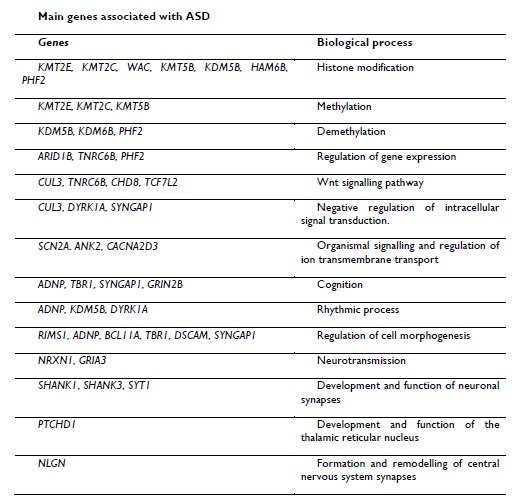
Research has been done to try to understand the genetic mechanism by which ASD is generated, however, the main obstacle is that ASD manifests a spectrum of symptoms that are caused by multiple genes, and has been shown to be largely caused as a result of alterations in genes that are involved in brain development.8 Although several genes have been identified as ASD genes, including NRXN1, SHANK1, SHANK3 and PTCHD1, much remains to be known about the extent of the underlying mutations.12 Table 1 shows the main genes associated with ASD.13 The genetic contribution to ASD is therefore heterogeneous, with many loci involved in the development of the disorder at different levels in different individuals,14 which is the reason why the phenotypic expression of these genes is highly variable in each person. Currently, it has not been possible to identify a common locus related to susceptibility to ASD. In addition to this, epigenetic mechanisms have recently been related as potential contributors to the development of ASD, that is to say, environmental factors have an important role in the predisposition to develop these disorders.15
Table 1. Genes that have been associated with ASD and the biological processes that are carried out (Taken and modified from Ramaswami and Geschwind, 2018).
The miRNAs are small non-coding RNA molecules, between 18 to 25 nucleotides, whose main target is messenger RNA (mRNA), which they degrade or inhibit, resulting in a decrease or inhibition in the synthesis of the resulting protein; and they are located in the genome, even in areas without a known function.6-7,16 The miRNAs work as post-transcriptional regulators, by binding to mRNA in the 3´ untranslated region, resulting in the inhibition of mRNA.17 Generally, more than 2000 miRNAs have been determined in humans and about 50 % of human genes are predicted to be regulated by them, and furthermore, they are involved in all the important functions of the cell: proliferation, differentiation, cell cycle control, DNA repair, apoptosis, immunity, stress, homeostasis, aging, development and metabolism.1,17-19
Moreover, miRNAs play a very important role in the development of the Central Nervous System (CNS), carrying out important functions in neurogenesis, synaptogenesis and neuronal migration.20 Therefore, it is not surprising that the dysregulation of miRNAs leads to changes in behaviour and cognition, observed in many neurodegenerative disorders such as Alzheimer’s Disease (AD), Parkinson´s Disease (PD), Amyotrophic Lateral Sclerosis (ALS), polyglutamine (polyQ) disorders such as Huntington´s disease (HD) and spinocerebellar ataxias (SCAs) and ASD.20-22
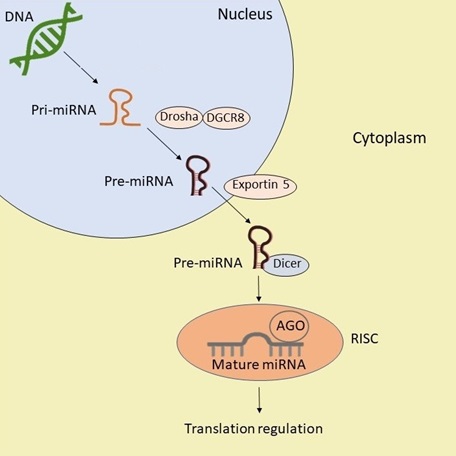
The miRNAs biogenesis is initiated in the cell nucleus with the generation of a primary transcript (pri-miRNA) by the RNA polymerase II and its subsequent conversion of pri-miRNA into pre-miRNA which is arbitrated by the microprocessor complex Drosha/DGCR8.1,23 The pre-miRNA is transported to the cytoplasm by the exportin-5 where is processed by the RNase Ⅲ enzyme Dicer and the mature miRNA is generated (a double stranded miRNA). Mature miRNA together with the Argonaute (AGO) protein form the RNA-induced silencing complex (RISC).24 Is in this complex where mature miRNA is guided to complementary sequences in 3’ or 5’ where it regulates the translation of target mRNAS (figure 1).1,23,25
One possible mechanism by which miRNAs regulate gene expression is by inducing the binding and activity of RNA polymerase in specific genes, although, they may also be available to bind to chromatin modifiers, resulting in transcriptional silencing, working as regulators of epigenetic controllers or as regulator of the specific expression of genes.26-27 In addition to their function as part of the epigenetic machinery, they can also be modified by this same mechanism, changing their expression under both, physiological and pathological conditions, and there is evidence that the expression of miRNAs varies in healthy and disease state.28
The key of the miRNAs functioning is the formation of the RISC, composed of AGO protein and a miRNA molecule.29-30 This complex can downregulate gene expression by two posttranscriptional mechanism: cleavage of mRNA or translational repression.2 miRNAs will recognize their mRNA target through the complementarity base pairing in the 3’ or 5’ untranslated region.24 Depending on the base pairing level (complete or partial) the result could be degradation of the mRNA or translational repression.31 Although the most studied function of the miRNAs is the downregulation of gene expression, there is evidence that indicate that some miRNAs could enhance the gene expression, however the mechanism is not well known.32
Figure 1. Scheme of miRNAs Biogenesis and function. After being transcribed in the nucleus, the pri-miRNA is processed by Drosha/DGCR8 into pre-miRNA which is transported to the cytoplasm by exportin 5, where is processed by Dicer, generating mature miRNA, it is this mature miRNA that exerts its function of translation regulation.
Initially, miRNAs were identified in cellular microenvironments.33 However, they have also been found in extracellular microenvironments, including those derived from blood sources such as serum and plasma, and other fluids such as tears, urine, saliva, amniotic fluid, peritoneal fluid, cerebrospinal fluid, seminal fluid, synovial fluid, pleural fluid, bronchial lavage, follicular fluid, colostrum and breast milk, that is to say, in practically all the fluids of the body.33-34 Therefore, miRNAs can be considered as potential biomarkers for the diagnosis of various diseases, including ASD.
Control of miRNA-dependent protein synthesis is an important posttranscriptional regulator in almost all aspects of nervous system development.6 Many miRNAs are regulated during Central Nervous System (CNS) development and expressed in adult brain indicating their important roles in neural development and function.35 Depending on the target gene, the same miRNA can promote or inhibit developmental processes.23 Because miRNAs are also subject to epigenetic modifications, their function can change depending on time and space, so the same miRNA can be involved in different decisions about the cell fate or development of cells.26,36 miRNAs have been found to play important roles at different stages of brain function, particularly in neuronal plasticity and neuronal development.,37 Furthermore, recent studies have shown that miRNAs may be one of the possible causes associated with the development of ASD, mainly in mouse models, due to the observation of the correlation between the loss of function of some miRNAs and an altered social behaviour and that the upregulation of others favoured the dendritic growth.38-40
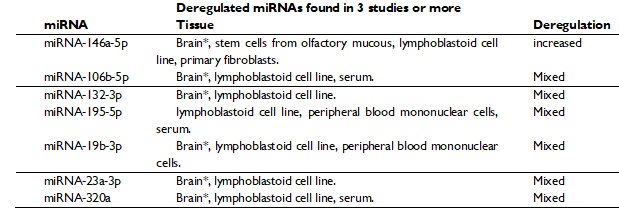
In addition to the above, have been reported altered expressions of miRNAs in whole blood, serum, lymphoblastic lines and post-mortem brains of individuals with ASD, compared to healthy individuals.8 156 altered miRNAs have been reported in patients with ASD, of which 7 were found in at least 3 different studies (table 2).17
Among the miRNAs found in different studies miRNA-146a-5p is the miRNA that has the most evidence in relation to ASD, since it was found to be significantly increased in 3 studies and in 4 different types of tissue; brain, olfactory mucosal stem cells, lymphoblastoid cell lines and primary lymphoblasts.41-43 Some target genes for this miRNA are NOTCH1, NLGN, GRIA3, SYT1, among others, all these genes are essential for the development and function of the brain.14,16
Using a microarray assay a miRNA, miRNA-486-3p, was found to be altered in patients with ASD. This miRNA is specific to the brain with possible roles in neuronal differentiation in humans, the study showed increased expression in patients with ASD in whole blood.44
An outstanding miRNA is miRNA-132, that promotes the growth and maturation of neurons in the adult hippocampus and could be essential in the regulation of cognition, learning and memory formation.45-46 It participates in the regulation of neuronal differentiation, maturation and function, axon growth, neural migration and plasticity.47 Disruption of this miRNA results in, among other things, aberrant neural development.45 Of the multiple mRNAs that are targeted by miRNA-132, two of them, MeCP2 and the activation protein P250GAP have been studied and found to have fundamental roles in the development and function of dendritic spines. An animal model study of mice showed that prenatal exposure to valproic acid, used in animal models of ASD, increased the levels of miRNA-132 in the brain of male and female embryos.48 This may assume that miRNA-132 levels are increased in alterations of neuronal activity in the embryonic period, at least in ASD-like behavioural animal models produced by prenatal exposure to valproic acid.
Another interesting miRNA is miRNA-155p5 which is involved in several neuro-inflammatory diseases, including individuals with ASD who present neuro-inflammation due to microglial activation. Using quantitative real time PCR (QRT-PCR) in post-mortem human brain tissues this miRNA showed increased gene expression in the amygdala of children with ASD, indicating the presence of localised inflammation in the brain.49
Through animal models using KO mice or rats treated with valproic acid (VPA) it has been found that other miRNAs are associated with ASD, for example: a) miRNA-125b and miRNA-132 have opposite effects on dendritic spine morphology and contribute to the alteration of synaptic plasticity, regulating the expression of the FMR1 and UBE3A genes, respectively.50 b) in the brain of patients with ASD, miRNA-134-5p is overexpressed and is also upregulated in animals of the VPA models, this miRNA controls the development of the dendritic spines.51 c) miRNA-138-5p is abundant in the brain, is localized within dendrites and participates in the changes in the development of the dendritic spines and in the narrowing of the spine through the expression of the APT1 gene.52 d) A positive regulation of miRNA-34a and reduction in BCL2 expression levels were observed immediately after birth in rats treated with VPA in the cerebral cortex. Therefore, the miRNA-34a/BCL2 signalling pathway may be an activation mechanism for ASD.53 e) in the amygdala of patients with ASD, miRNA-181c and miRNA-30d are upregulated,54 the function of this miRNA is associated with the development of the nervous system; f) in rats treated with VPA, the miRNA-30d is positively regulated and its function is associated with the development of the nervous system.38
In reference to human studies, one of them analysed the expression of 125 mature miRNAs in the serum from 55 individuals with ASD and 55 control subjects using QRT-PCR, miRNAs were identified as differentially expressed in ASD individuals compare to the control, 8 were downregulated; miRNA-151a-3p, miRNA-181b-5p, miRNA-320a, miRNA-328, miRNA-433, miRNA-489, miRNA-572, and miRNA-663a, while five of them were upregulated miRNA-101-3p, miRNA-106b-5p, miRNA-130a-3p, miRNA-195-5p, and miRNA-19b-3p. Five of this miRNAs showed hight predictive power for differentiate individuals with ASD.37 Another study in saliva from 24 individuals with ASD showed fourteen miRNAs differentially dysregulated compared to controls, of which 9 were upregulated; miRNA- 7-5p, miRNA-28-5p, miRNA-127-3p, miRNA-140-3p, miRNA-191-5p, miRNA-218-5p, miRNA-335-3p, miRNA-628-5p, miRNA-2467-5p, and miRNA-3529-3p, whereas miRNA-23a-3p, miRNA-27a-3p, miRNA-30e-5p, and miRNA-32-5p were downregulated.55
Table 2. miRNAs reported as deregulated in ASD in 3 or more studies (modified from Fregeac et al, 2016).
6. miRNAs as biomarkers and their therapeutic potential in ASD
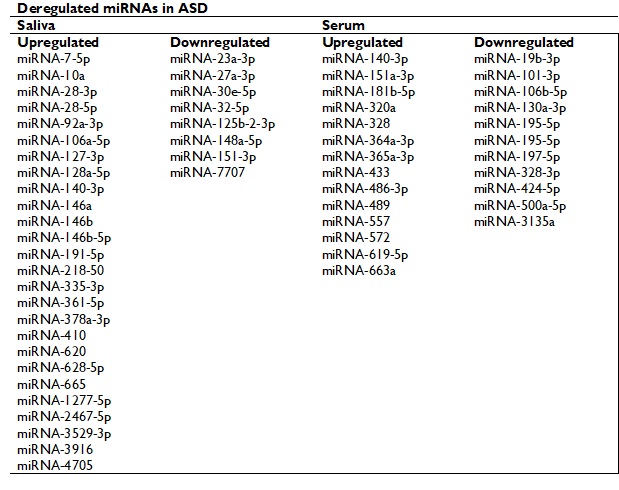
The ASD can be detected as young as 18 months of age based on developmental history and behaviour, without any medical test available today.56 The use of miRNAs as biomarkers has been extensively investigated, and due to their stability and ease of detection in small amounts of different biofluids such as blood and even serum, and also because several of them can be detected simultaneously, they could be a very useful tool for predicting ASD in the first months of life, even before behavioural signs can be detected. Furthermore, the development of tests to predict ASD by detecting miRNAs using biofluids (saliva or serum) is in the early stages, where certain limitations must be considered: the methodology and those that result from the lack of functional consequences of the miRNAs.33 Another limitation is that many of these miRNAs are involved and participate in multiple neurodegenerative processes, which implies a difficulty in differentiating ASD from some other neural development disorders. Even so, miRNAs remain a promising option to become a non-invasive screening test for ASD, which would allow detecting the disease at an earlier stage, even in the first two years of life, enabling therapeutic intervention in earlier stages. In this sense, several miRNAs have been described as dysregulated in the serum and saliva of individuals with ASD, and that could be used as promising biomarkers for the early detection of the disorder (table 3).33,57-58
Finally, because miRNAs may play a role in the development of ASD and because several of them can be detected in biofluid such as saliva and serum, this makes miRNAs eligible as candidates to be evaluated for detection and as therapeutic targets for ASD. Furthermore, most of the sequences of mature human miRNAs are 100 % identical to rodents, allowing their use as models for preclinical evaluations of efficacy, toxicity and pharmacodynamics.59-60
Table 3. miRNAs that have been described as dysregulated in the serum and saliva of individuals with ASD.
Studies indicate that several miRNAs are directly related to the regulation of the brain development and are directly involved to ASD, mainly in animal models. Other studies have attempt to approach to the diagnostic use with serum and saliva samples from individuals with ASD finding promising results. Due to these observations, miRNAs have the potential to be used as biomarkers for the diagnosis and prognosis of ASD.
In this way advantages and disadvantages must be considered, the miRNAs could be found in body fluids as saliva or serum, they are stable, and several can be detected at the same time. The main limitation is that the miRNAs participate in the nervous system development and therefore in multiple neurodegenerative processes, which implies a difficulty in differentiating ASD from some other neural development disorders.
Currently, ASD has the capacity to become a serious public health problem, and due to the etiopathological complexity of the disorder, its investigation and treatment are difficult, coupled with the fact that the tools used in the diagnose of this condition basically consist in a series of test, questionnaires and behaviour observations done to the subject that will measured the 3 affected domains of the ASD, but sometimes the underdiagnosis could disparage the real condition of the subject.
Is this complexity that makes miRNAs ideal candidates for use in the early diagnosis and treatment of the disorder. Although few miRNAs have been consistently identified as dysregulated in ASD by different studies, the evidence suggests that they contribute to its development. There are still several questions to which research effort must be devoted to answer them, since as miRNAs are epigenetically influenced mechanisms, it must be clarified how they are differentially expressed during the development of the disorder and which cells are the ones where they exert their effect. Although these limitations do not detract from current advances that point to the use of miRNAs as biomarkers, in order to consider them in practice as possible treatments for ASD, preclinical studies are required in which the key could be to discover how to specifically carry these miRNAs to the brain, which would open the doors for clinical use.
The authors declare no conflicts of interests.
Supported by a research grant from Consejo Nacional de Ciencia y Tecnología (Conacyt), with research grant number 467819.
1. Christodoulatos GS, Dalamaga M. Micro-RNAs as clinical biomarkers and therapeutic targets in breast cancer: Quo vadis? World J Clin Oncol 2014 5: 71–81.
2. Bartel DP. MicroRNAs: Genomics, Biogenesis, Mechanism, and Function Review. Cell 2004 116: 281–97.
3. Arberas C, Ruggieri V. Autismo y epigenética. Un modelo de explicación para la comprensión de la génesis en los trastornos del espectro autista. Med (B Aires) 2013 73: 20–9.
4. Manzo-Denes J. Un segundo espectro del autismo: de la conducta a la neurona A second spectrum for autism: from behavior to the neuron. 2019 10: 1–20.
5. Rogel-Ortiz FJ. Autismo. Gac Med Mex 2005 141: 143–7.
6. Rajman M, Schratt G. microRNAs in neural development: from master regulators to fine-tuners. Development 2017 144: 2310–22.
7. Lau NC, Lim LP, Weistein EG, Bartel DP. An abundant class of tiny RNAs with probably regulatory roles in Caenorhabditis elegans. Science 2001 294: 858–62.
8. Duarte F V, Palmeira CM, Rolo AP. The emerging role of MitomiRs in the pathophysiology of human disease. En: Santulli G. microRNA: Medical Evidence, Springer 2015: 123-154.
9. Maenner MJ, Shaw KA, Baio J, Washington A, Patrick M, DiRienzo M. Prevalence of autism spectrum disorder among children aged 8 years - Autism and developmental disabilities monitoring network, 11 Sites, United States, 2016. MMWR Surveill Summ 2020 69: 1–12.
10. Fombonne E, Marcin C, Manero AC, Bruno R, Diaz C, Villalobos M, Ramsay K, Nealy B. Prevalence of Autism Spectrum Disorders in Guanajuato, Mexico: The Leon survey. J Autism Dev Disord 2016 46: 1669–85.
11. Stewart J, Vigil D. Refining best practices for the diagnosis of autism: A comparison between individual healthcare practitioner diagnosis and transdisciplinary assessment. Nevada J Public Heal 2014 11: 1–12.
12. Woodbury-Smith M, Scherer SW. Progress in the genetics of autism spectrum disorder. Dev Med Child Neurol 2018 60: 445–51.
13. Ramaswami G, Geschwind DH. Genetics of autism spectrum disorder. En: Geschwind DH PAulson HL, Klein C. Handbook of Clinical Neurology. Elsevier B.V 2018: 321–329.
14. Persico AM, Napolioni V. Autism genetics. Behav Brain Res 2013 251: 95–112.
15. Meek SE, Lemery-Chalfant K, Jahromi LB, Valiente C. A review of gene–environment correlations and their implications for autism: A conceptual model. Psychol Rev 2013 120: 497–521.
16. Aqeilan RI, Calin GA, Croce CM. MiR-15a and miR-16-1 in cancer: Discovery, function and future perspectives. Cell Death Differ 2010 17: 215–20.
17. 17. Fregeac J, Colleaux L, Nguyen LS. The emerging roles of MicroRNAs in autism spectrum disorders. Neurosci Biobehav Rev 2016 71: 729–38.
18. Blenkiron C, Miska EA. miRNAs in cancer : approaches , aetiology , diagnostics and therapy. 2007 16: 106–13.
19. Kozomara A, Griffiths-Jones S. MiRBase: Integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res 2011 39: 152–7
20. Sehovic E, Spahic L, Smajlovic-Skenderagic L, Pistoljevic N, Dzanko E, Hajdarpasic A. Identification of developmental disorders including autism spectrum disorder using salivary miRNAs in children from Bosnia and Herzegovina. PLoS One 2020 15: 1–18.
21. Abe M, Bonini NM. MicroRNAs and neurodegeneration: Role and impact. Trends Cell Biol 2013 23: 30–6.
22. Chan AWS, Kocerha J. The path to microRNA therapeutics in psychiatric and neurodegenerative disorders. Front Genet 2012 3: 1–10.
23. Ruiz R, Garrido E, Velázquez-flores MÁ. Nuevos e inesperados mecanismos de biogénesis y acción de los microRNAs. 2016 35: 55–70.
24. Bushati N, Cohen SM. MicroRNA functions. Annu Rev Cell Dev Biol 2007 23: 175–205.
25. Bartel DP. MicroRNAs: Target Recognition and Regulatory Functions. Cell 2009 136: 215–33.
26. Piletič K, Kunej T. MicroRNA epigenetic signatures in human disease. Arch Toxicol 2016 90: 2405–19.
27. Fabbri M, Calore F, Paone A, Galli R, Calin GA. Epigenetic Regulation of miRNAs in Cancer. Epigenetic Alterations Oncog 2012 754: 137–48.
28. Van den Hove DL, Kompotis K, Lardenoije R, Kenis G, Mill J, Steinbusch HW, Lesch P, Fitzsimons CP, De Strooper B, Rutten BPF. Epigenetically regulated microRNAs in Alzheimer ’ s disease. Neurobiol aging 2014 35: 731–45.
29. Catalanotto C, Cogoni C, Zardo G. MicroRNA in Control of Gene Expression : An Overview of Nuclear Functions. 2016 17.
30. Liu J, Carmell MA, Rivas F V, Marsden CG, Michael J. Argonaute2 is the catalytic engine of mammalian RNAi. Science 2004 305: 1437–41.
31. He L, Hannon GJ. MicroRNAs : small RNAs with a big role in gene regulation. Nat Rev Genet 2004 5: 522–31.
32. Place RF, Li L, Pookot D, Noonan EJ, Dahiya R. MicroRNA-373 induces expression of genes with. Proc Natl Acad Sci 2018 115: 1608–13
33. Salloum-Asfar S, Satheesh NJ, Abdulla SA. Circulating miRNAs, Small but Promising Biomarkers for Autism Spectrum Disorder. Front Mol Neurosci 2019 12: 1–10.
34. Weber JA, Baxter DH, Zhang S, Huang DY, Huang KH, Lee MJ, et al. HHS Public Access. 2016 56: 1733–41.
35. Cao D, Li L, Chan W. MicroRNAs : Key Regulators in the Central Nervous System and Their Implication in Neurological Diseases. Int J Mol Sci 2016 17: 1–28.
36. Faravelli I, Corti S. MicroRNA-Directed Neuronal Reprogramming as a Therapeutic Strategy for Neurological Diseases. Mol Neurobiol 2018 55: 4428–36.
37. Mundalil Vasu M, Anitha A, Thanseem I, Suzuki K, Yamada K, Takahashi T, et al. Serum microRNA profiles in children with autism. Mol Autism 2014 5: 1–9.
38. Schepici G, Cavalli E, Bramanti P, Mazzon E. Autism spectrum disorder and mirna: An overview of experimental models. Brain Sci 2019 9: 1–16.
39. Valluy J, Bicker S, Aksoy-aksel A, Lackinger M. A coding-independent function of an alternative Ube3a transcript during neuronal development. Nat Neurosci 2015 18: 666–73.
40. Lackinger M, Sungur AÖ, Daswani R, Soutschek M, Bicker S, Stemmler L, et al. A placental mammal-specific microRNA cluster acts as a natural brake for sociability in mice. EMBO Rep 2019 20: 1–11.
41. Mor M, Nardone S, Sams DS, Elliott E. Hypomethylation of miR-142 promoter and upregulation of microRNAs that target the oxytocin receptor gene in the autism prefrontal cortex. Mol Autism 2015 6: 1–11.
42. Nguyen LS, Lepleux M, Makhlouf M, Martin C, Fregeac J, Siquier-Pernet K, et al. Profiling olfactory stem cells from living patients identifies miRNAs relevant for autism pathophysiology. Mol Autism 2016 7: 1–13.
43. Talebizadeh Z, Butler MG, Theodoro MF. Feasibility and relevance of examining lymphoblastoid cell lines to study role of microRNAs in autism. Autism Res 2008 1: 240–50.
44. Popov NT, Madjirova NP, Minkov IN. Micro RNA HSA-486-3P Gene Expression Profiling in the Whole Blood of Patients with Autism. Biotechnol Biotechnol Equip 2012 26: 3385–8.
45. Hansen KF, Karelina K, Sakamoto K, Wayman GA, Impey S, Obrietan K. miRNA-132: A dynamic regulator of cognitive capacity. Brain Struct Funct 2013 218: 817–31.
46. Vaishnavi V, Manikandan M, Tiwary BK, Munirajan AK. Insights on the Functional Impact of MicroRNAs Present in Autism-Associated Copy Number Variants. PLoS One 2013 8: 1–13.
47. Qian Y, Song J, Ouyang Y, Han Q, Chen W, Zhao X, et al. Advances in roles of miR-132 in the nervous system. Front Pharmacol 2017 8: 1–9.
48. Hara Y, Ago Y, Takano E, Hasebe S, Nakazawa T, Hashimoto H, et al. Prenatal exposure to valproic acid increases miR-132 levels in the mouse embryonic brain. Mol Autism 2017 8: 1–9.
49. Almehmadi KA, Tsilioni I, Theoharides TC. SHORT REPORT Increased Expression of miR-155p5 in Amygdala of Children With Autism Spectrum Disorder. 2019 00:1–6.
50. Dieter Edbauer, Neilson JR, Foster KA, Wang C-F, Seeburg DP, Batterton MN, et al. Regulation of synaptic structure and function by FMRP- associated. Neuron 2010 65: 373–84.
51. Hirsch MM, Deckmann I, Fontes-Dutra M, Bauer-Negrini G, Della-Flora Nunes G, Nunes W, et al. Behavioral alterations in autism model induced by valproic acid and translational analysis of circulating microRNA. Food Chem Toxicol 2018 115: 336–43.
52. Siegel G, Obernosterer G, Fiore R, Oehmen M, Christensen M, Khudayberdiev S, et al. A functional screen implicates microRNA-138-dependent regulation of the depalmitoylation enzyme APT1 in dendritic spine morphogenesis Gabriele. Nat Cell Biol 2013 11: 705–16.
53. Dai X, Yin Y, Qin L. Valproic acid exposure decreases the mRNA stability of Bcl-2 via up-regulating miR-34a in the cerebellum of rat. Neurosci Lett 2017 657: 159–65.
54. Olde Loohuis NFM, Kole K, Glennon JC, Karel P, Borg G Van der, Gemert Y Van, et al. Elevated microRNA-181c and microRNA-30d levels in the enlarged amygdala of the valproic acid rat model of autism. Neurobiol Dis 2015 80: 42–53.
55. Hicks SD, Ignacio C, Gentile K, Middleton FA. Salivary miRNA profiles identify children with autism spectrum disorder, correlate with adaptive behavior, and implicate ASD candidate genes involved in neurodevelopment. BMC Pediatr 2016 16: 1–11.
56. Hyman SL, Levy SE, Myers SM, Children ON, Disabilities W. Identification , Evaluation , and Management of Children With Autism Spectrum Disorder. Pediatrics 2020 145.
57. Narayanan R, Schratt G. miRNA regulation of social and anxiety ‑ related behaviour. Cell Mol Life Sci 2020.
58. Ozkul Y, Taheri S, Bayram KK, Sener EF, Ecmel M, Öztop DB, et al. A heritable profile of six miRNAs in autistic patients and mouse models. Sci Rep 2020 10.
59. Fregeac J, Colleaux L, Nguyen LS. The emerging roles of MicroRNAs in Autism Spectrum Disorders. Neurosci Biobehav Rev 2016 71: 730–38.
60. Rooij E van, Kauppinen S. Development of microRNA therapeutics is coming of age. EMBO Mol Med 2014 6: 851–64.
| Recibido: 09 de julio, 2020 | Aceptado: 20 de diciembre, 2020 |
*Correspondencia en: Fausto Rojas Durán, Instituto de Investigaciones Cerebrales, Universidad Veracruzana, Xalapa, Ver. México. Avenida Dr. Luis Castelazo Ayala S/N, colonia Industrial Ánimas, C.P. 91190. Tel.: (228) 8418900 Ext. 13051. E-mail: frojas@uv.mx
Este es un artículo de libre acceso distribuido bajo los términos de la licencia de Creative Commons, (http://creamasal@unam.mxtivecommons.org/licenses/by-nc/3.0), que permite el uso no comercial, distribución y reproducción en algún medio, siempre que la obra original sea debidamente citada.